Rebecca Page, Principal Investigator
Department of Molecular Biology, Cell Biology and Biochemistry
My laboratory uses X-ray crystallography , NMR spectroscopy and small angle X-ray scattering (SAXS) in combination with biochemistry and genetics to understand the molecular basis of protein function, with a particular interest in understanding how bacterial signaling proteins regulate biolfim formation and antibiotic resistance, how targeting proteins direct the activity of ser/thr phosphatases and how dual specificity and tyrosine phosphatases regulate MAPK function.
Biofilms, persistence and antibiotic resistance
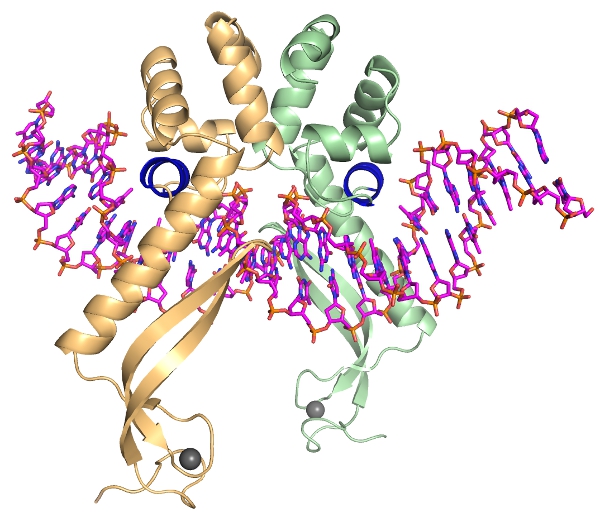 Bacterial biofilms are complex communities of cells containing an increased prevalence of dormant cells known as persisters, which underlie the multidrug tolerance of biofilms. However, a detailed understanding of how they assemble and how they are regulated at a molecular level is still only rudimentarily understood.
Using a combination of structural, genetic and biochemical studies, we discovered that the E. coli protein mqsR, which is the most highly upregulated gene in persister cells, together with mqsA, are the defining members of an entirely new family of toxin:antitoxin systems which are essential for persistence. We are now invesitgating how the MqsA:MqsR TA system regulates biofilm formation and antibiotic resistance in vivo and new chemical tools to inhibit mqsRA mediated persistence in the cell.
Bacterial biofilms are complex communities of cells containing an increased prevalence of dormant cells known as persisters, which underlie the multidrug tolerance of biofilms. However, a detailed understanding of how they assemble and how they are regulated at a molecular level is still only rudimentarily understood.
Using a combination of structural, genetic and biochemical studies, we discovered that the E. coli protein mqsR, which is the most highly upregulated gene in persister cells, together with mqsA, are the defining members of an entirely new family of toxin:antitoxin systems which are essential for persistence. We are now invesitgating how the MqsA:MqsR TA system regulates biofilm formation and antibiotic resistance in vivo and new chemical tools to inhibit mqsRA mediated persistence in the cell.
PP1 holoenzymes in the nucleus
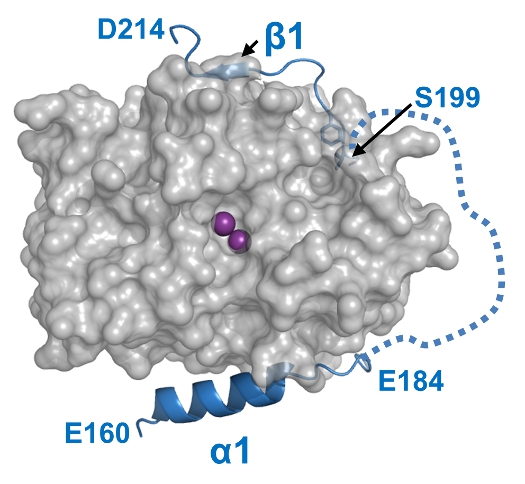 The regulation of the Ser/Thr phosphatase protein phosphatase 1 (PP1) is controlled by a diverse array of regulatory proteins. However, how these proteins direct the specificity of PP1 is not well understood. In addition, while 100’s of cell biology and biochemical reports describe key biological roles for PP1, very few structural efforts have so far been successful. We are focusing on two PP1-targeting proteins-- the nuclear ihibitor of PP1 (NIPP1) and the PP1 nuclear targeting subunit (PNUTS)--to elucidate, at a molecular level, the biological functions and modes of action of these key nuclear PP1 holoenzymes in order to provide unique novel insights into the molecular regulation of PP1.
The regulation of the Ser/Thr phosphatase protein phosphatase 1 (PP1) is controlled by a diverse array of regulatory proteins. However, how these proteins direct the specificity of PP1 is not well understood. In addition, while 100’s of cell biology and biochemical reports describe key biological roles for PP1, very few structural efforts have so far been successful. We are focusing on two PP1-targeting proteins-- the nuclear ihibitor of PP1 (NIPP1) and the PP1 nuclear targeting subunit (PNUTS)--to elucidate, at a molecular level, the biological functions and modes of action of these key nuclear PP1 holoenzymes in order to provide unique novel insights into the molecular regulation of PP1.
Regulation of MAPKs by phosphatases
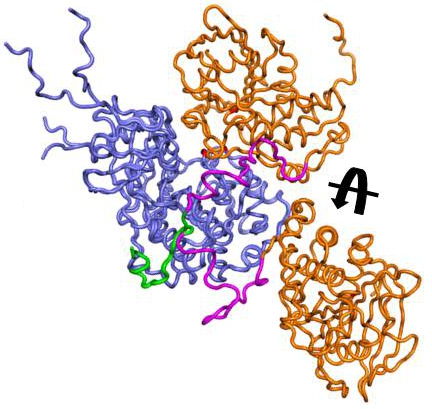 MAP kinases transduce environmental and developmental signals into adaptive and programmed responses. In the cell, MAPK are regulated by numerous phosphatases. For example, the
hematopoietic tyrosine phosphatase (HePTP) negatively
regulates T cell activation in lymphocytes via ERK2 dephosphorylation.
However, only very limited structural information
is available for these biologically important complexes. We are combining X-ray crystallography, NMR spectroscopy, SAXS to characterize the structures of the MAPK:PTP complexes and ITC to understand the energetics of binding.
MAP kinases transduce environmental and developmental signals into adaptive and programmed responses. In the cell, MAPK are regulated by numerous phosphatases. For example, the
hematopoietic tyrosine phosphatase (HePTP) negatively
regulates T cell activation in lymphocytes via ERK2 dephosphorylation.
However, only very limited structural information
is available for these biologically important complexes. We are combining X-ray crystallography, NMR spectroscopy, SAXS to characterize the structures of the MAPK:PTP complexes and ITC to understand the energetics of binding.
Graduate Student Dana Lord recieved her PhD (5/2014). Congratulations Dr. Lord!
Undergraduate Robin Rolander was awarded the prestigous Brown Univeristy UTRA award (4/2014).
Graduate Student Dana Lord received a travel award and was also invited to give a lecture at the 2013 American Crystallographic Association meeting in Hawaii (8/2013).
Our review, the 'Structural Basis of PP1 Regulation and Specificity' is the FEBS Journal most downloaded paper in 2013.
Outreach: The 5th annual Protein Science Workshop and Lecture, a collaboration between the Page lab and the Biotechnology Certificate Program at CCRI was held.
For a full publication list, click here.
Choy et al (2014) Understanding the antagonism of retinoblastoma protein dephosphorylation by PNUTS provides insights into the PP1 regulatory code. PNAS 111: 4097-4102.
Peti & Page (2013) Molecular Basis of MAPK Regulation Protein Science, 22: 1698-1710.
Wolfgang Peti, Ph.D., Brown University.
Lutz Tautz, Ph.D., The Burnham Institute.
Thomas Wood, Ph.D, Pennsylvania State University.